RQC, UPS and cytoskeletal dynamics during erythroid differentiation/megakaryopoiesis
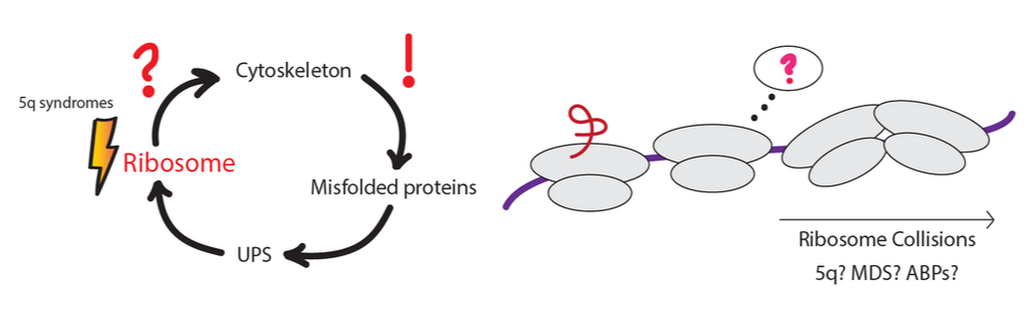
Only a few E3 ligases have been characterized and known to play significant roles in erythropoiesis/megakaryopoiesis. Even though majority of protein turnover during differentiation can be attributed to programmed cell death (protease activated), there remain a slew of proteasome dependent mechanisms which might be critical for terminal erythroid differentiation as well as platelet biogenesis. Datasets from various studies have revealed that E3 ligases and dUBs are involved in proteostasis during erythroid differentiation. On the other hand, mono-ubiquitination events on ribosomal proteins via E3 ligases are well known mechanisms to circumvent ribosomal stalling. We know that 5q syndromes and Diamond Blackfan Anemia have inherent tendencies to display stalled ribosomes. Are there E3 ligases involved in ribosomal ubiquitylation (RRub) we can pinpoint to be a part of this process but have become dysfunctional during disease? Additionally, recent evidence has shown that the UPS can regulate the actin cytoskeleton. Thus, will RRub affect erythroid differentiation or megakaryopoiesis via regulation of the actin cytoskeleton?
Research interests in this area will rely on the role ribosomal ubiquitylation plays during erythroid differentiation and megakaryopoiesis as well as whether destabilization of multi-ordered complexes can instigate the proteasome, E3 ligases & dUBs in blood disorders and congenital bone marrow failures.
Only a few E3 ligases have been characterized and known to play significant roles in erythropoiesis/megakaryopoiesis. Even though majority of protein turnover during differentiation can be attributed to programmed cell death (protease activated), there remain a slew of proteasome dependent mechanisms which might be critical for terminal erythroid differentiation as well as platelet biogenesis. Datasets from various studies have revealed that E3 ligases and dUBs are involved in proteostasis during erythroid differentiation. On the other hand, mono-ubiquitination events on ribosomal proteins via E3 ligases are well known mechanisms to circumvent ribosomal stalling. We know that 5q syndromes and Diamond Blackfan Anemia have inherent tendencies to display stalled ribosomes. Are there E3 ligases involved in ribosomal ubiquitylation (RRub) we can pinpoint to be a part of this process but have become dysfunctional during disease? Additionally, recent evidence has shown that the UPS can regulate the actin cytoskeleton. Thus, will RRub affect erythroid differentiation or megakaryopoiesis via regulation of the actin cytoskeleton?
Research interests in this area will rely on the role ribosomal ubiquitylation plays during erythroid differentiation and megakaryopoiesis as well as whether destabilization of multi-ordered complexes can instigate the proteasome, E3 ligases & dUBs in blood disorders and congenital bone marrow failures.
Focal adhesions involved in erythropoiesis/megakaryopoiesis and platelet biogenesis
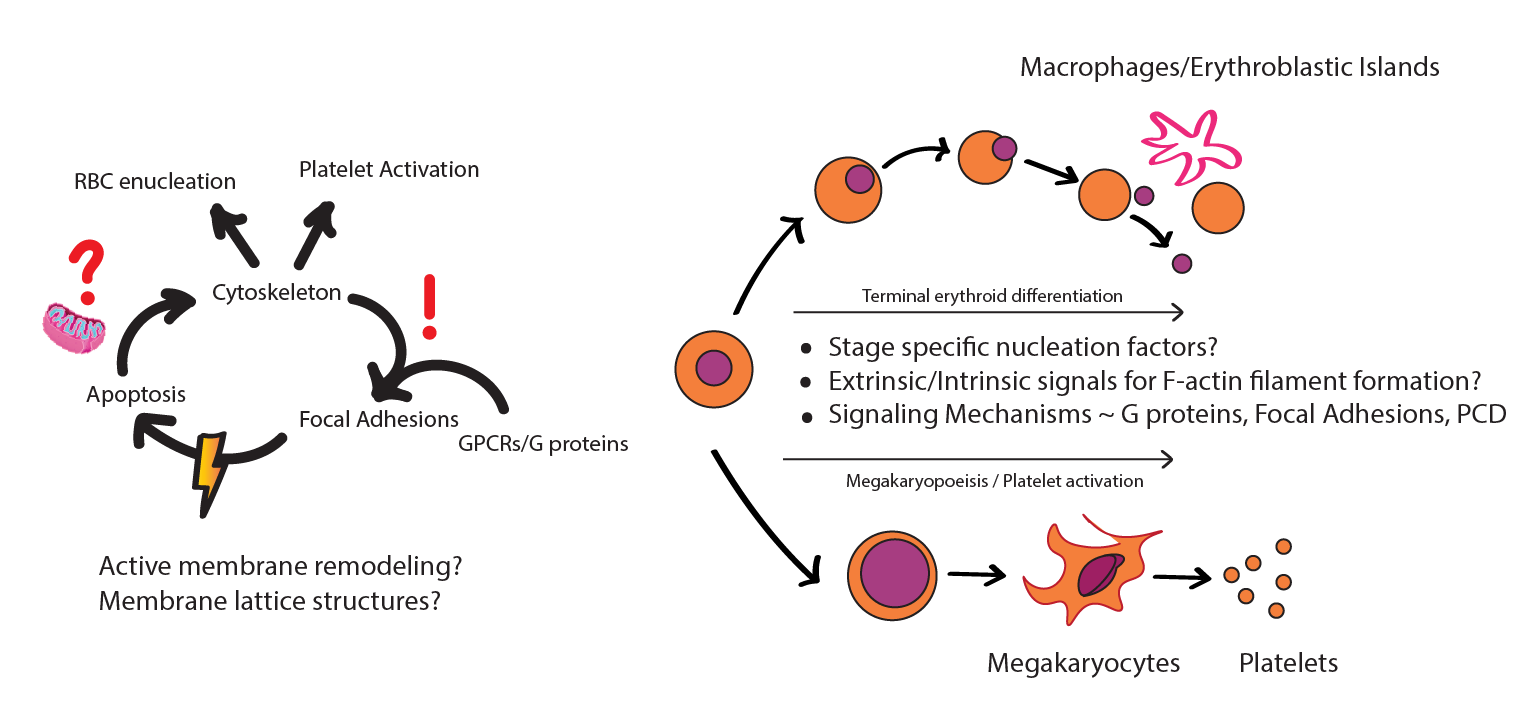
Hematopoietic stem cells while going into lineage commitment into erythroid pool will co-regulate adhesion and integrin molecules to prevent red cell clumping and agglutination during development. On the other hand, focal adhesion kinases are involved in megakaryocytic maturation, platelet biogenesis and platelet activation. Our recent findings have suggested that certain focal adhesion molecules are important in erythroid differentiation. Do these adhesion molecules render inherent mechanical complexities to the red cell membrane which facilitates erythrocyte detachment from the ECM and in erythroblastic islands? Preliminary data has also suggested that loss of certain FAs leads to higher propensity of clots and lowered platelet counts in mice. How will FAs regulate cell membrane lattices, F-actin organization and
Research interests in this area will focus on uncovering the molecular aspects of the orchestra between focal adhesions (FA) and integrins as they play out during erythroid differentiation. We will look into how FAs will regulate erythroid differentiation and investigate molecular changes during blood disorders such as sickle cell anemia, hemolytic anemias and spherocytosis.
Hematopoietic stem cells while going into lineage commitment into erythroid pool will co-regulate adhesion and integrin molecules to prevent red cell clumping and agglutination during development. On the other hand, focal adhesion kinases are involved in megakaryocytic maturation, platelet biogenesis and platelet activation. Our recent findings have suggested that certain focal adhesion molecules are important in erythroid differentiation. Do these adhesion molecules render inherent mechanical complexities to the red cell membrane which facilitates erythrocyte detachment from the ECM and in erythroblastic islands? Preliminary data has also suggested that loss of certain FAs leads to higher propensity of clots and lowered platelet counts in mice. How will FAs regulate cell membrane lattices, F-actin organization and
Research interests in this area will focus on uncovering the molecular aspects of the orchestra between focal adhesions (FA) and integrins as they play out during erythroid differentiation. We will look into how FAs will regulate erythroid differentiation and investigate molecular changes during blood disorders such as sickle cell anemia, hemolytic anemias and spherocytosis.
Small molecule/ CRISPR screens to find novel players in erythropoiesis, megakaryopoiesis and platelet biogenesis
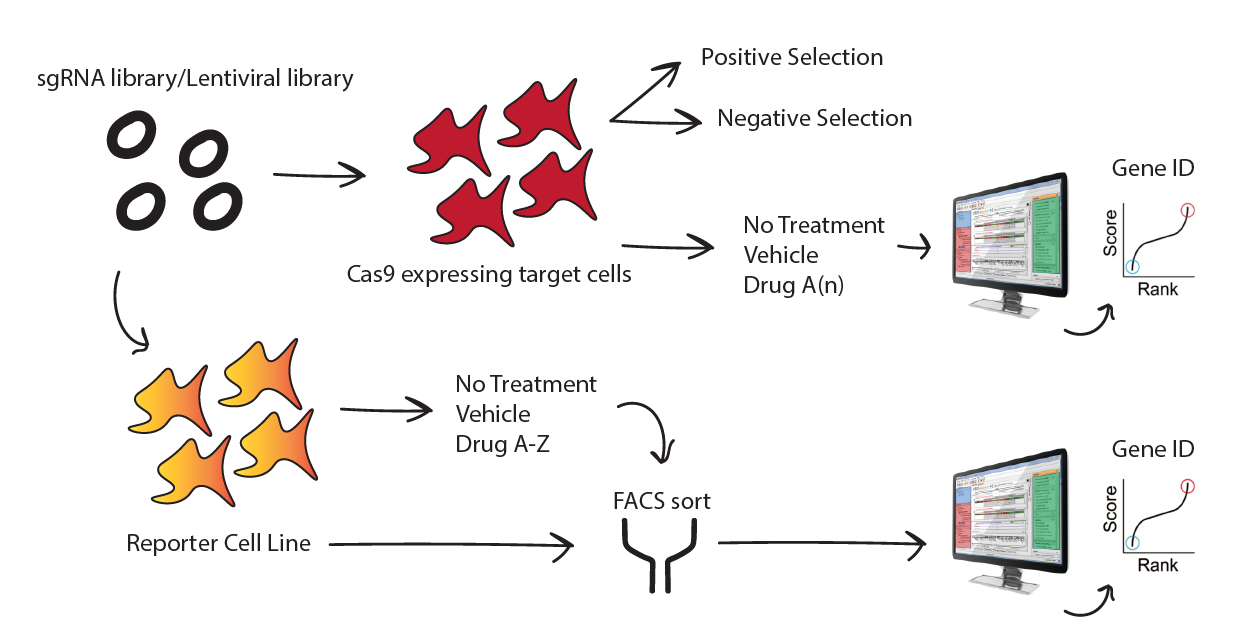
A long-standing goal for my research has been to execute small molecule screens, utilize chemical libraries and develop CRISPR libraries for various UPS (E3 ligases & dUBs), translational (RPS) and signaling molecules (G-proteins, GPCRs, focal adhesions) in erythroid and megakaryocytic lineages. For erythroid cells we want to look for small molecules or protein families/proteins which disrupt hemoglobin accumulation, surface marker expression (GPA, CD71, CD47 etc) or even enucleation. In MEPs and megakaryoblasts we want to look for similar effects perturbing CD41, MPL, CD42b etc. To this end current tech such as scRNA-seq combined with high content imaging would be highly useful to look for molecular and physiological effects post exposure of primary (CD34+) cells to chemical / CRISPR libraries during CEP/CMP transitions.
Research interests in this area will focus on discovery aspects of HSC biology with emphasis on screening for defects in any given lineage from a common myeloid progenitor. The implications of this research will directly benefit clinical applications and possibly even instigate IND studies for various diseases such as thrombocytopenia, leukemias, anemias and MDS.
A long-standing goal for my research has been to execute small molecule screens, utilize chemical libraries and develop CRISPR libraries for various UPS (E3 ligases & dUBs), translational (RPS) and signaling molecules (G-proteins, GPCRs, focal adhesions) in erythroid and megakaryocytic lineages. For erythroid cells we want to look for small molecules or protein families/proteins which disrupt hemoglobin accumulation, surface marker expression (GPA, CD71, CD47 etc) or even enucleation. In MEPs and megakaryoblasts we want to look for similar effects perturbing CD41, MPL, CD42b etc. To this end current tech such as scRNA-seq combined with high content imaging would be highly useful to look for molecular and physiological effects post exposure of primary (CD34+) cells to chemical / CRISPR libraries during CEP/CMP transitions.
Research interests in this area will focus on discovery aspects of HSC biology with emphasis on screening for defects in any given lineage from a common myeloid progenitor. The implications of this research will directly benefit clinical applications and possibly even instigate IND studies for various diseases such as thrombocytopenia, leukemias, anemias and MDS.